Sequence databases with taxonomy annotations
See also
Microbial taxonomy
The SILVA and Greengenes "taxonomies"
Taxonomic nomenclatures (naming systems)
Taxonomy errors in 16S databases
Conflicts between 16S trees and taxonomy
Specialized 16S rRNA sequence databases providing
taxonomy annotations include Greengenes, LTP, RDP, and SILVA.
At the time of writing (mid-July 2018), it appears that
Greengenes may be defunct as the last release was v13.5 from May 2014.
LTP (Living Tree Project) is based on sequences of type strains and
isolates classified on the basis of observed traits. The other databases are
much larger, containing mostly environmental sequences.
In the RDP database, taxonomy annotations of
environmental sequences are predicted by the Naive
Bayesian Classifier (NBC). I estimate the error rate of these
annotations to be roughly 10%. A subset of type strain and isolate sequences
is provided as training data for the Classifier.
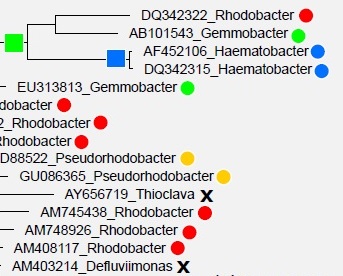 Greengenes
and SILVA are annotated by a combination of database-specific computational
prediction methods and manual curation based on predicted phylogenetic trees
inferred from multiple sequence alignments. Roughly
one in five of these annotations are wrong,
almost certainly because of branching
order errors in the trees.
Greengenes
and SILVA are annotated by a combination of database-specific computational
prediction methods and manual curation based on predicted phylogenetic trees
inferred from multiple sequence alignments. Roughly
one in five of these annotations are wrong,
almost certainly because of branching
order errors in the trees.