distmx_split_identity command
The distmx_split_identity command divides sequences into subsets such that the top-hit identity is a given value. This is used to create test-training pairs for cross-validation by identity . Input is a distance matrix created by the calc_distmx command.
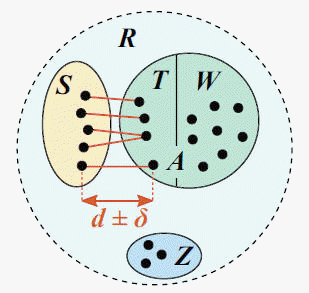 The range of allowed top-hit identities is given by the -mindist and -maxdist options, which must both be specified. Identities are specified as distances as they appear in the matrix. For example, -mindist 0.025 -maxdist 0.035 specifies an identity range from 97.5% to 96.5%.
The range of allowed top-hit identities is given by the -mindist and -maxdist options, which must both be specified. Identities are specified as distances as they appear in the matrix. For example, -mindist 0.025 -maxdist 0.035 specifies an identity range from 97.5% to 96.5%.
Output is a tabbed text given by the -tabbedout option. Fields are:
#1 Subset name.
#2 Label1
#3 Label2
#4 Dist
There are four subsets with names 1, 2, 1x and 2x. Label1 is the label of a sequence in the subset given by #1. Label2 is the top hit in the other subset (1 or 2), and Dist is the distance between Label1 and Label2.
Subsets 1 and 2 have top hits to each other in the specified range.
Subset 1x has lower identities with subset 2, and can therefore be added to the training set if subset 2 is the query set. Similarly, subset 2x has lower identities with subset 1 and can be added to the training set if subset 1 is the query.
Example
usearch -calc_distmx tax_16s.fa -maxdist 0.2 -termdist 0.3 -tabbedout distmx_16s.txt
usearch -distmx_split_identity distmx_16s.txt -mindist 0.025 -maxdist 0.035 \
-tabbedout subsets.txt

