Viral RdRp and its sequence analysis challenges
Viral RdRp protein
Viral RNA dependent RNA polymerase (RdRp) is the protein responsible for copying the genome of an RNA virus.
Almost all RNA viruses make an RdRp protein
(
Wikipedia article on RdRp).
Exceptions include RNA retroviruses
(
retroviruses), where the polymerase
protein is a
reverse transcriptase, and satellite viruses
such as
Hepatitis Delta Virus which require RdRp
from another virus (the so-called helper virus) to be present to support replication.
RdRp proteins are not named RdRp
Typically, the protein with RdRp function is not officially called RdRp. For example, in nidoviruses the RdRp protein is called Nsp9
or
Nsp12, and in the
lambda 3 phage it is simply called the lambda protein
(see PDB
1MUK).
This complicates database searches by gene name.
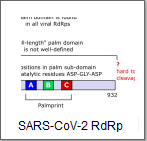

Where does RdRp sequence terminate?
Sometimes, RdRp has a simple coding sequence (
CDS) beginning with a START codon
and ending at a STOP codon. In such cases, it is straightforward to identify RdRp as the nucleotide or translated amino acid
sequence of this CDS (assuming of course that you know the genetic code of the host). However, things are not always that simple.
RdRp often occurs in a longer ORF which codes for multiple proteins;
for example in SARS-CoV-2 RdRp function is found in
Nsp12,
one of several non-structural proteins coded in the long ORF1ab (21,555nt)
which accounts for most of the genome (29,903nt). ORF1ab is translated into a long peptide (pp1ab), which is then
split into mature proteins by a
cleavage enzyme.
In at least one family (narnaviruses), RdRp is not a single protein; it is assembled as a complex from
preptides coded in multiple
genome segments (https://pubmed.ncbi.nlm.nih.gov/34688782/).
Cleavage sites are hard to find
Unlike START and STOP codons, cleavage sites are not easily recognized by sequence analysis, and in
fact are not known in many fully-sequenced genomes
(
https://pubmed.ncbi.nlm.nih.gov/27567259/.
Therefore, the beginning and end of the RdRp sequence in a genome or metagenomic contig may be
difficult to identify. In practice, this problem means that predicted RdRp sequences are often
truncated or trimmed to shorter subsequences such as a partial domain.
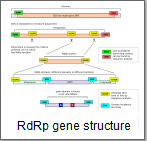
RdRp domains
The RdRp protein is usually constructed from two or more domains.
The term
domain generally means a segment
which folds independently and may be found in combination with other domains in different proteins,
but the definition is not precise and may be used in different ways in different contexts.
Domain boundaries are fuzzy
It can be clear that some residues are in a particular domain
(for example, the [G/S]DD motif is the palm domain).
However, there are no sharp boundaries where two adjacent residues are definitively in
different domains. Thus, in contrast to CDS which has clear boundaries (start
/ stop codons), domain boundaries are fuzzy.
Palm domain
The palm domain is found in all known viral RdRps
(
Jia and Gong 2019,
te Velthuis 2014).
Other named domains are usually present in an RdRp protein; for example, nidoviruses such as SARS-CoV-2 have
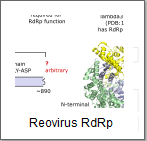
a NiRAN domain at the N terminal before the palm, and reoviruses have an
unnamed N-terminal domain before the palm and a bracelet
domain after the palm. Regions outside of the palm domain are generally not well understood,
and it is often not known whether they are essential for RdRp function or perform
some other function.
Varying domains in RdRp
As the examples of nidoviruses (NiRAN+palm) and reoviruses (N-terminal+palm+bracelet) illustrate,
the domain content of RdRp varies in different families, and except for the palm domain
are often not well characterised. This adds to the difficulty of identifying the boundaries
of the RdRp coding sequence in genomes which are far diverged from well-characeterized viruses,
and similarly in metagenomic contigs.
Permuted domains
Several families have permuted RdRp genes
(
Sabanadzovic2009,
Ambrose2009,
Ferrero2021,
Gorbalenya2002)
where the palm domain is permuted.
